| Drug Design Laboratory |
|
 |
| Sitemap |
| Print Version |
| Login |
| About CMSimple |
| News > Research projects > Homology modeling |
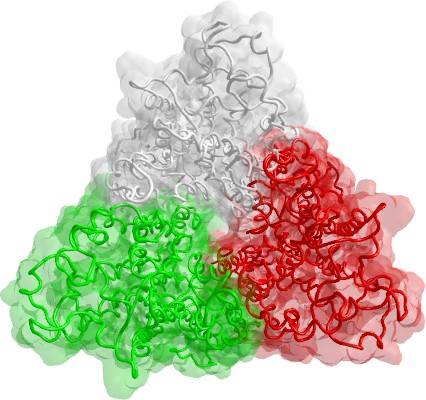
Homology modeling
In these last years, the number of experimentally resolved transmembrane proteins is considerably growing providing useful templates to model many other proteins. In particular, the availability of the experimental crystal structure of bovine rhodopsin has strongly supported the homology modelling of full-length G-protein coupled receptors (GPCR) and, in recent years, several reliable GPCR homology models appeared in literature, showing that they can be successfully used for virtual screening and ligand optimization. However, the rhodopsin based homology models have two main drawbacks:
Hence, it comes as no surprise that some rhodopsin-independent methods have been proposed in last years. In general, a recent trend in folding prediction is to favour the local homology combining more predictive algorithms (the so-called meta-prediction, as implemented in the ShotGun technology). In principle, these approaches (1) divide the aminoacidic sequence in fragments, (2) predict the folding of each fragment using different methods, (3) exhaustively combine the predicted fragments obtaining several models, and (4) select the best model using suitable score functions. The GPCR modelling well suits this fragmental strategy because (1) the aminoacidic sequence is clearly divisible in 15 structural fragments (namely, 7 transmembrane helices, 6 loops, and 2 terminal segments), (2) the fragment prediction is not really blind, but it is well known that the transmembrane segments assume helix conformations and the loops must have a global “U” shape in which the loop ends are close enough to join the adjacent TM segments, and (3) the rhodopsin crystal structure can be still exploited as template to drive the final assembly of predicted fragments. In our reserch laboratory, we modeled some intersting proteins using the fragmental approach:
All these models are available on request.
|
| < | Privacy and cookies | > |
